Research Scope
This repository combines three related threads of chest X-ray work:
- TB classification with TorchXRayVision and PyTorch Lightning
- lung segmentation with MONAI and PyTorch Lightning
- diffusion-based image generation on cropped chest X-rays
The current emphasis is the segmentation study.
Core Question
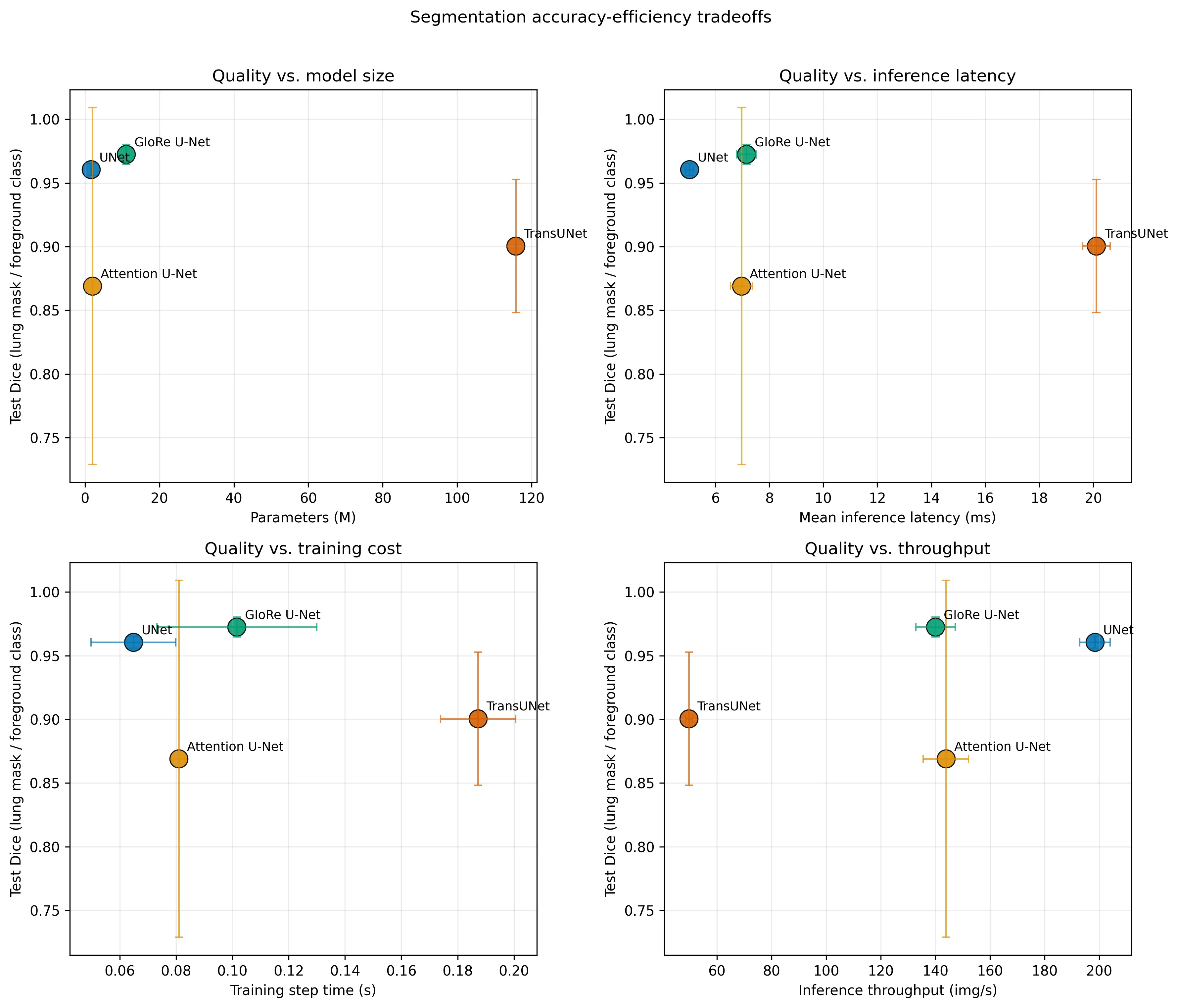
The central comparison asks how much segmentation quality is gained from stronger global-context modeling, and what that costs in model size, training time, and inference latency.
Compared Models
- U-Net
- Attention U-Net
- GloRe U-Net
- TransUNet
Findings
The current multi-seed study described in the README suggests:
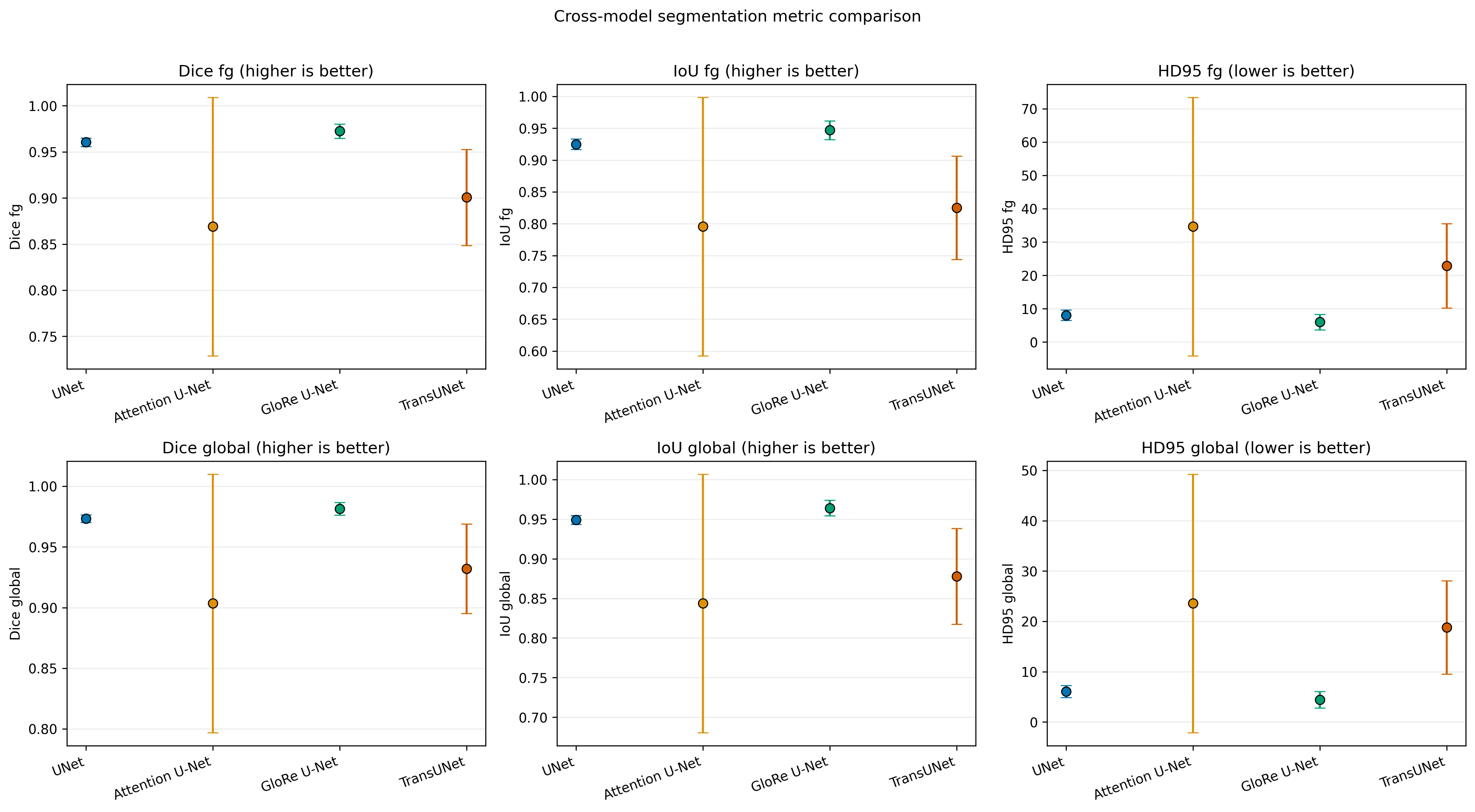
- baseline U-Net and GloRe U-Net are the strongest overall performers
- GloRe U-Net slightly improves foreground Dice while staying in a similar latency range
- Attention U-Net is more variable across seeds
- TransUNet is heavier and slower in this setup without clearly beating the strongest CNN-based models on average
Why It Matters
This page belongs under Research because the value is in controlled comparison, experiment infrastructure, and report-ready outputs. It is a good example of model evaluation work that treats quality and efficiency as linked decisions rather than separate concerns.